Note
Go to the end to download the full example code.
Domain-wide Gridded Emission Scaling¶
Apply a constant factor to a subset of emission variables in a file.
The basic steps are:
Copy a file.
Select species to scale NOx species.
Scale by a multiplicative factor.
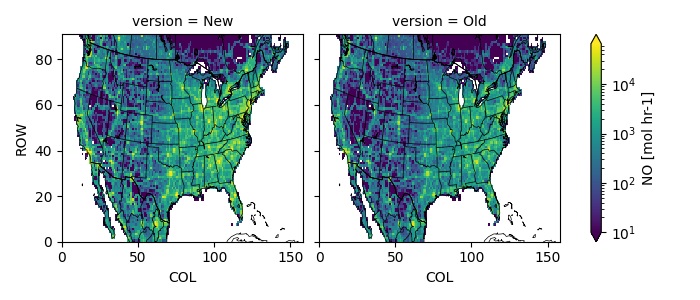
Plot the old and new result.
Reminder: You must have already activated your python environment.
Configuration¶
# date to process
date = '20160610'
# file to copy
oldpath = f'../../camx/emiss/camx_area.mobile.{date}.36km.nc'
# new file to create
newpath = f'outputs/camx_area_2x.mobile.{date}.36km.nc'
# species to scale
scalekeys = ['NO', 'NO2', 'HONO'] # list of strings or set to 'all'
# factor to scale by
factor = 2.
# path for figure comparing
figpath = 'outputs/scaled_gridemiss_scalewholedomain.png'
Imports and File Prep¶
import netCDF4
import shutil
import os
os.makedirs('outputs', exist_ok=True)
shutil.copyfile(oldpath, newpath)
'outputs/camx_area_2x.mobile.20160610.36km.nc'
Open Files¶
Open the existing file in read-only mode
Open the new file in editable append mode
infile = netCDF4.Dataset(oldpath, mode='r')
outfile = netCDF4.Dataset(newpath, mode='a')
Select Emission Variables to Scale¶
if scalekeys is all, identify all emission variables by unit
if scalekeys == 'all':
scalekeys = [
k for k, v in outfile.variables.items()
if v.units.strip() in ('mol hr-1', 'g hr-1')
]
noscalekeys = sorted(set(outfile.variables).difference(scalekeys))
print('INFO:: no scale', noscalekeys)
print('INFO:: to scale', scalekeys)
INFO:: no scale ['ACET', 'ALD2', 'ALDX', 'BENZ', 'BNZA', 'CH4', 'CO', 'CPRM', 'ECH4', 'ETFLAG', 'ETH', 'ETHA', 'ETHY', 'ETOH', 'FORM', 'FPRM', 'IOLE', 'ISOP', 'ISP', 'IVOA', 'KET', 'MEOH', 'NA', 'NH3', 'NVOL', 'OLE', 'PAL', 'PAR', 'PCA', 'PCL', 'PEC', 'PFE', 'PH2O', 'PK', 'PMG', 'PMN', 'PNH4', 'PNO3', 'POA', 'PRPA', 'PSI', 'PSO4', 'PTI', 'SO2', 'SULF', 'TERP', 'TFLAG', 'TOL', 'TOLA', 'TRP', 'UNK', 'UNR', 'X', 'XYL', 'Y', 'latitude', 'longitude']
INFO:: to scale ['NO', 'NO2', 'HONO']
Apply Scaling¶
for skey in scalekeys:
print('INFO:: scaling', skey, 'by', factor)
outvar = outfile.variables[skey]
invar = infile[skey]
outvar[:] = factor * invar[:]
outfile.sync()
del outfile
INFO:: scaling NO by 2.0
INFO:: scaling NO2 by 2.0
INFO:: scaling HONO by 2.0
Plot Comparison¶
import pyrsig
import pycno
import matplotlib.colors as mc
oldfile = pyrsig.open_ioapi(oldpath)
newfile = pyrsig.open_ioapi(newpath)
key = scalekeys[0]
compfile = newfile[[]]
compfile['New'] = newfile[key].sum('LAY').mean('TSTEP')
compfile['Old'] = oldfile[key].sum('LAY').mean('TSTEP')
Z = compfile.to_dataarray(dim='version')
Z.attrs.update(oldfile[key].attrs)
fca = Z.plot(col='version', norm=mc.LogNorm(10, compfile['Old'].max()))
pycno.cno(oldfile.crs_proj4).drawstates(ax=fca.axs)
fca.fig.savefig(figpath)
Total running time of the script: (0 minutes 8.584 seconds)