Note
Go to the end to download the full example code.
Modify a Selected Source¶
Point sources may be facilities or individual stacks. This example demonstrates how to apply a scaling factor to all sources in some region. The region can be as simple as a rectangular box, a circle, or any geographic shape you can define.
The basic steps are:
Copy an existing emissions file.
Define an area within which to apply emissions sclaing.
Define a multiplicative scaling factor
Apply the scaling factor to all cells within the area.
Make a plot demonstrating the change.
Reminder: You must have already activated your python environment.
Configuration¶
Find a source in the in “Online 2020 NEI Data Retrieval Tool” https://www.epa.gov/air-emissions-inventories/2020-national-emissions-inventory-nei-data - As an example, I chose single stack with the largest NO rate; then updated
the lon/lat to the nearest source in the NEI 2020 inventory
You’ll have a chance to adjust your shape until you capture the intended source.
# date to process
date = '20160610'
# file to copy
oldpath = f'../../camx/ptsrce/point.camx.ptnonipm.{date}.nc'
# new file to create
newpath = f'outputs/point.camx.ptnonipm_edit.{date}.nc'
# figure demonstrating change
figpath = 'outputs/example_ptsrce_scaleregion.png'
# Region to modify
shape = 'box' # box, circle, or custom
center = (-82.5436, 36.5224)
dx = 250 # m (used as radius for circle)
dy = 250 # m (ignored for circle)
# factor to apply within the shape
scalekeys = ['NO', 'NO2', 'HONO'] # a list of species or 'all'
factor = 1.2
Imports and Folders¶
import numpy as np
import netCDF4
import pyproj
import shutil
from shapely import points, box
import os
Define Output and Remove if it exists¶
Use an existing file as a template
Create a copy for direct modification
os.makedirs('outputs', exist_ok=True)
if os.path.exists(newpath):
os.remove(newpath)
shutil.copyfile(oldpath, newpath)
'outputs/point.camx.ptnonipm_edit.20160610.nc'
Open Files¶
Open the existing file in read-only mode
Open the new file in append mode
oldf = netCDF4.Dataset(oldpath, mode='r')
newf = netCDF4.Dataset(newpath, mode='a')
Select Emission Variables to Scale¶
If scalekeys is all, select all emission variables by units
if scalekeys == 'all':
scalekeys = [
k for k, v in newf.variables.items()
if v.units.strip() in ('mol hr-1', 'g hr-1')
]
noscalekeys = sorted(set(newf.variables).difference(scalekeys))
print('INFO:: no scale', noscalekeys)
print('INFO:: to scale', scalekeys)
INFO:: no scale ['ACET', 'ALD2', 'ALDX', 'BENZ', 'BNZA', 'CH4', 'CL2', 'CO', 'CPRM', 'ECH4', 'ETFLAG', 'ETH', 'ETHA', 'ETHY', 'ETOH', 'FORM', 'FPRM', 'HCL', 'IOLE', 'ISOP', 'ISP', 'IVOA', 'KET', 'MEOH', 'NA', 'NH3', 'NVOL', 'OLE', 'PAL', 'PAR', 'PCA', 'PCL', 'PEC', 'PFE', 'PH2O', 'PK', 'PMG', 'PMN', 'PNH4', 'PNO3', 'POA', 'PRPA', 'PSI', 'PSO4', 'PTI', 'SO2', 'SULF', 'TERP', 'TFLAG', 'TOL', 'TOLA', 'TRP', 'UNR', 'XYL', 'flowrate', 'pigflag', 'plume_bottom', 'plume_top', 'plumerise', 'saoverride', 'stkdiam', 'stkheight', 'stkspeed', 'stktemp', 'xcoord', 'ycoord']
INFO:: to scale ['NO', 'NO2', 'HONO']
Define relevant sources¶
Create a projection
- Define an area to scale in
in a rectangle,
in a circle, or
within a complex custom shape
attrs = {k: oldf.getncattr(k) for k in oldf.ncattrs()}
proj4tmpl = '+proj=lcc +lat_0={YCENT} +lon_0={P_GAM}'
proj4tmpl += ' +lat_1={P_ALP} +lat_2={P_BET} +R=6370000 +units=m +no_defs'
proj4str = proj4tmpl.format(**attrs)
proj = pyproj.Proj(proj4str)
cx, cy = proj(*center) # project to x/y
if shape == 'box':
myshp = box(cx - dx, cy - dy, cx + dx, cy + dy) # - define a rectangle
elif shape == 'circle':
myshp = points(cx, cy).buffer(dx) # - define a circle
elif shape == 'custom':
# define some custom shape based on a shapefile
import geopandas as gpd
# downloaded from census.gov
cntypath = '../../www2.census.gov/geo/tiger/TIGER2020/COUNTY/tl_2020_us_county.zip'
# Read just in the continental US
cnty = gpd.read_file(cntypath, bbox=(-135, 20, -60, 60))
# Select counties in Georgia, reproject to grid space, collapse to one big shape (i.e, state)
# myshp = cnty.query('STATEFP == "13"').to_crs(proj.srs).union_all()
# Select DeKalb county in Georgia, reproject to grid space, collapse to one shape
myshp = cnty.query('GEOID == "13091"').to_crs(proj.srs).union_all()
else:
raise KeyError(f'For shape, got {shape} -- use box or circle')
psxy = points(oldf['xcoord'][:], oldf['ycoord']) # make points for each source
ismine = myshp.intersects(psxy) # find all points in shape
print(f'INFO:: {ismine.sum()} point sources in myshp')
INFO:: 60 point sources in myshp
Point Source Distances¶
Group stacks by unique distance from myshp perimeter.
- Report all clusters within maxdist.
Check that the number inside (0m) matches your expectation.
Check for nearby clusters that might be the same facilty.
Consider adjusting dx, dy or shape type to better capture the source.
maxdist = 10e3 # stop looking at 10km
psdist = points(cx, cy).distance(psxy)
fdists, fcnts = np.unique(psdist, return_counts=True)
print('INFO:: Stacks by distance from myshp center')
for fdist, fcnt in zip(fdists, fcnts):
print(f'INFO:: {fdist:6.0f}m: {fcnt}')
if fdist > maxdist:
break
INFO:: Stacks by distance from myshp center
INFO:: 223m: 4
INFO:: 224m: 56
INFO:: 1016m: 2
INFO:: 1321m: 6
INFO:: 3564m: 1
INFO:: 3624m: 14
INFO:: 9100m: 14
INFO:: 19298m: 24
Apply Scaling¶
for skey in scalekeys:
print(f'INFO:: multiplying {skey} by {factor:.1%} at selected point sources')
newvals = oldf[skey][:].copy()
newvals[:, ismine] *= factor
newf[skey][:] = newvals
newf.sync()
del newf
INFO:: multiplying NO by 120.0% at selected point sources
INFO:: multiplying NO2 by 120.0% at selected point sources
INFO:: multiplying HONO by 120.0% at selected point sources
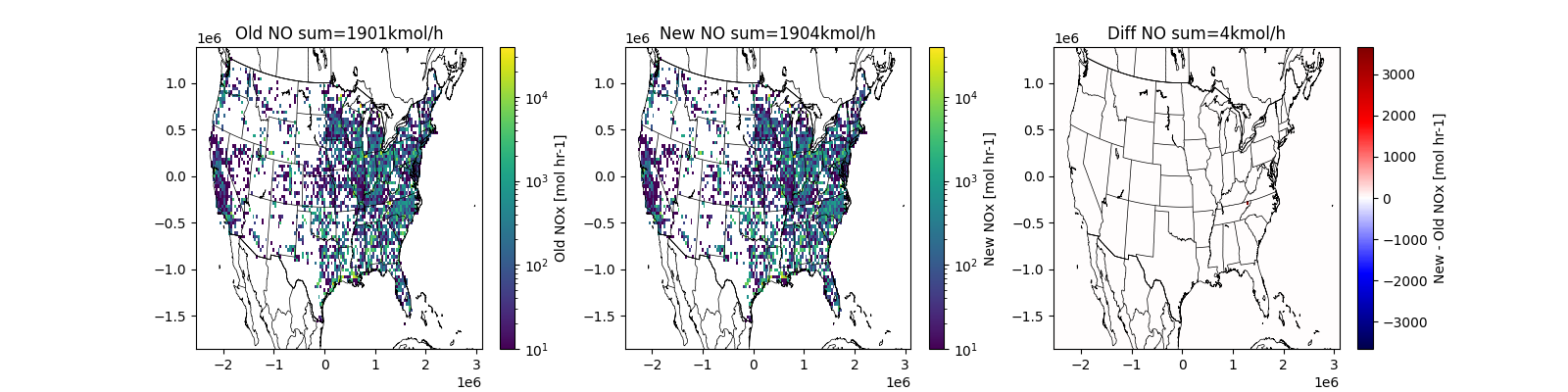
Visualize Source to confirm operational¶
Create gridded data for quick visualiation
Show new, original and difference.
import matplotlib.colors as mc
import matplotlib.pyplot as plt
import pycno
import numpy as np
oldf = netCDF4.Dataset(oldpath, mode='r')
newf = netCDF4.Dataset(newpath, mode='r')
x = oldf['xcoord'][:]
y = oldf['ycoord'][:]
# add all emission keys
zold = sum([oldf[ek][:].mean(0) for ek in scalekeys])
znew = sum([newf[ek][:].mean(0) for ek in scalekeys])
xedges = np.arange(0.5, oldf.NCOLS) * oldf.XCELL + oldf.XORIG
yedges = np.arange(0.5, oldf.NROWS) * oldf.YCELL + oldf.YORIG
Hold, _, _ = np.histogram2d(y, x, weights=zold, bins=[yedges, xedges])
Hnew, _, _ = np.histogram2d(y, x, weights=znew, bins=[yedges, xedges])
gskw = dict(left=0.333, right=0.967)
fig, axx = plt.subplots(1, 3, figsize=(16, 4))
opts = dict(norm=mc.LogNorm(10), cmap='viridis')
qm = axx[0].pcolormesh(xedges, yedges, Hold, **opts)
fig.colorbar(qm, label='Old NOx [mol hr-1]')
axx[0].set(title=f'Old NO sum={Hold.sum() / 1e3:.0f}kmol/h')
qm = axx[1].pcolormesh(xedges, yedges, Hnew, **opts)
fig.colorbar(qm, label='New NOx [mol hr-1]')
axx[1].set(title=f'New NO sum={Hnew.sum() / 1e3:.0f}kmol/h')
opts['norm'] = mc.TwoSlopeNorm(0)
opts['cmap'] = 'seismic'
qm = axx[2].pcolormesh(xedges, yedges, Hnew - Hold, **opts)
fig.colorbar(qm, label='New - Old NOx [mol hr-1]')
axx[2].set(title=f'Diff NO sum={(Hnew.sum() - Hold.sum()) / 1e3:.0f}kmol/h')
pycno.cno(proj=proj).drawstates(resnum=1, ax=axx)
fig.savefig(figpath)
Extra Credit¶
Modify the custom section to scale all point sources in your home state.
Modify the custom section to scale all point sources in the counties in a nonattainment area.
Total running time of the script: (0 minutes 4.185 seconds)