Note
Go to the end to download the full example code.
Simple OMI¶
This example compares CAMx to the Ozone Monitoring Instrument (OMI) for NO2 tropospheric vertical column densitities (tropNO2). OMI’s tropNO2 is derived from a measurement along the path light from the sun and reflected off the surface of the earth. The original measurement is called a slant column density (SCD). The SCD is split into tropospheric and stratospheric parts, based on a model profile (a priori). The a priori and slant path are combined with a sensitivity vector (scattering weight) to convert the tropospheric SCD to tropNO2 [molecules/cm2]. CAMx can output mixing ratios at all levels, which can be converted to molecules/cm2 and summed to make a tropNO2 product. Note, however, that the CAMx tropNO2 inherently assumes a different model profile. A more complete comparison would adjust OMI to use CAMx’s profile, or adjust CAMx to use OMI’s profile. Each has strengths and weaknesses.
Reminder: You must have already activated your python environment.
Imports and Folders¶
import pyrsig
import os
os.makedirs('outputs', exist_ok=True)
Configuration¶
cdate = '20160610' # Using first available CAMx date
odate = '2017-06-10' # oddly Jun 10-11 of 2016 have no OMI over the US
# Use dates to find avrg file and 3d met file
cpath = f'../../camx/outputs/CAMx.v7.32.36.12.{cdate}.3D.avrg.grd01.nc'
# If you used camx.tar.gz for inputs, update the met folder to metss
mpath = f'../../camx/met/camx.3d.36km.{cdate}.nc'
# Define output paths.
outstem = f'outputs/OMINO2_CAMx.v7.32.36.12.{cdate}.3D.avrg.grd01'
outpath = outstem + '.nc'
tilepath = outstem + '_tile.png'
scatterpath = outstem + '_scatter.png'
Data Acquisition¶
# Open a CAMx 3D CONC file for NO2
cf = pyrsig.open_ioapi(cpath)
# Open a 3d met file (pressure, z, temperature, humiditiy) and match avrg file time
mkeys = ['pressure', 'z', 'temperature', 'humidity']
mf = pyrsig.open_ioapi(mpath)[mkeys].interp(TSTEP=cf.TSTEP)
# Process OMI for the CAMx grid
api = pyrsig.RsigApi(bbox=(-130, 20, -60, 60), workdir='outputs', grid_kw=cf.attrs)
omids = api.to_ioapi('omi.l2.omno2.ColumnAmountNO2Trop', bdate=odate)
Derive Conversion Factors¶
OMI tropospheric NO2 column (tropNO2) in molecules/cm2
CAMx average mixing ratios (X) in ppmV of dry-air by layer (k)
Need dry-air intensity (I) molecules/cm2 by layer
CAMx tropNO2 = $sum_k{X(k) * I(k)}$
# Calculate layer depths from interface heights
dz = mf['z'].copy()
dz[:, 1:] = mf['z'].diff('LAY').data
# Combine pressure, temperature, humidity, and dz to get dry-air intensity
molecules_per_cm2 = (
mf['pressure'] * 100 # mb -> Pa
/ 8.314 / mf['temperature'] # Pa -> moles/m3
* (1 - mf['humidity'] / 1e6) # moles/m3 -> moles_dry/m3
* dz * 6.022e23 / 1e4 # moles_dry/m3 -> molecules_dry/m2
)
vcdno2 = (cf['NO2'] * 1e-6 * molecules_per_cm2.data).sum('LAY', keepdims=True)
vcdno2.attrs.update(
long_name='CAMx_tropNO2', units='molecules/cm2',
var_desc='Simple sum of molecules'
)
Organizing for output¶
# Allowing OMI coordinates (time) to override CAMx.
omids['CAMx_tropNO2'] = vcdno2.dims, vcdno2.data, vcdno2.attrs
# Mask missing values from OMI.
outf = omids.where(omids['COLUMNAMOUNTNO2'] > -9e36)
# Make OMI naming more clear.
outf = outf.rename(COLUMNAMOUNTNO2='OMI_tropNO2')
outf['OMI_tropNO2'].attrs.update(long_name='OMI_tropNO2')
# Save to disk
pyrsig.cmaq.save_ioapi(outf.drop_vars('TFLAG'), outpath)
Visualize Results¶
import matplotlib.pyplot as plt
import matplotlib.colors as mc
import pyrsig
import pycno
import numpy as np
from scipy.stats import linregress
# Average potential overlapping overpasses
omids = pyrsig.open_ioapi(outpath).mean(('TSTEP', 'LAY'))
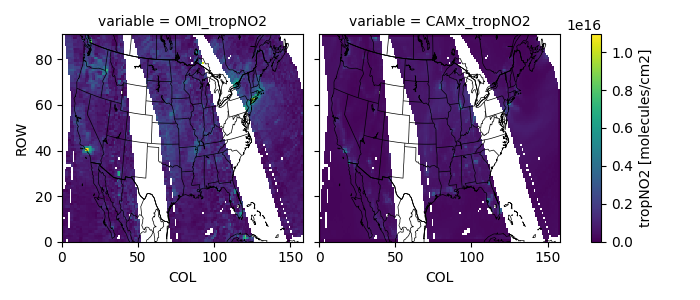
Make a tile-plot¶
Z = omids[['OMI_tropNO2', 'CAMx_tropNO2']].to_dataarray()
Z.attrs.update(long_name='tropNO2', units='molecules/cm2')
fca = Z.plot(col='variable')
pycno.cno(omids.crs_proj4).drawstates(ax=fca.axs)
fca.fig.savefig(tilepath)
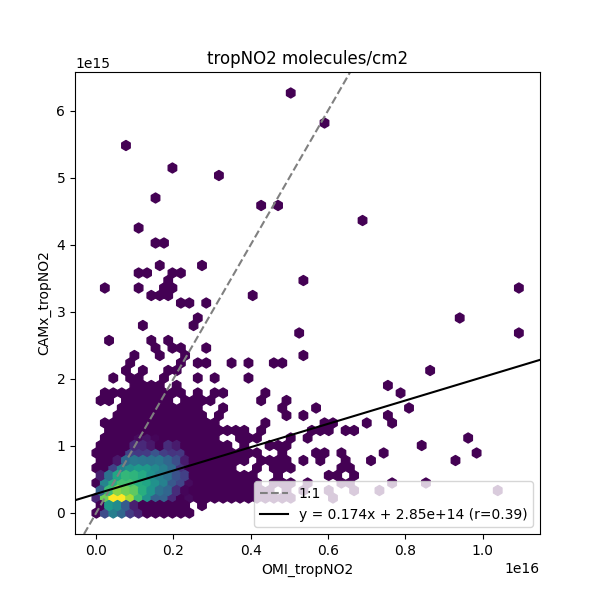
Make a scatter-plot¶
# Get values from OMI and CAMx separately
omivals = omids['OMI_tropNO2'].to_numpy().ravel()
camxvals = omids['CAMx_tropNO2'].to_numpy().ravel()
bothvalid = ~(np.isnan(omivals) | np.isnan(camxvals))
# Perform least-square regression
lr = linregress(omivals[bothvalid], camxvals[bothvalid])
lrlabel = f'y = {lr.slope:.3}x + {lr.intercept:.3} (r={lr.rvalue:.2f})'
# Create a hexbin plot
fig, ax = plt.subplots(figsize=(6, 6))
pc = ax.hexbin(x=omivals[bothvalid], y=camxvals[bothvalid], gridsize=50, norm=mc.LogNorm(20))
ax.set(xlabel='OMI_tropNO2', ylabel='CAMx_tropNO2', title='tropNO2 molecules/cm2')
ax.axline((0, 0), slope=1, label='1:1', color='grey', linestyle='--')
ax.axline((0, lr.intercept), slope=lr.slope, label=lrlabel, color='k')
ax.legend()
fig.savefig(scatterpath)
Total running time of the script: (0 minutes 12.816 seconds)